Resistotypes as a characterization of microbial communities associated with human health. Systematic Review
- Authors: Pavlenko A.V.1, Sonets I.V.1, Manolov A.I.1, Starikova E.V.1, Ilina E.N.1
-
Affiliations:
- Research Institute for Systems Biology and Medicine
- Issue: Vol 102, No 1 (2025)
- Pages: 112-126
- Section: REVIEWS
- URL: https://microbiol.crie.ru/jour/article/view/18666
- DOI: https://doi.org/10.36233/0372-9311-597
- EDN: https://elibrary.ru/zrutam
- ID: 18666
Cite item
Abstract
The concept of resistotype represents a new characterization of bacterial communities distinct from their taxonomic composition. The term “resistotype” is now commonly used to refer to specific clusters within a resistome that possess a characteristic phenotypic profile, or set of antimicrobial resistance genes. In contrast to traditional approaches that focus on individual resistance genes or phenotypic manifestations of resistance, resistome analysis allows resistance to be considered at a more comprehensive level, integrating different genes and their interactions within microbial populations or an entire microbial community.
The aims and objectives of the review are to analyze and summarize current data on how resistotypes of individual microbial species and bacterial communities are identified and analyzed.
Literature sources devoted to the identification of resistotypes of individual bacterial species and bacterial communities of humans and farm animals over the past 10 years were analyzed.
At the current moment, identification of microorganism resistotypes is not a common practice for studies related to resistance analysis. Phenotypic research methods, rarely supplemented by genetic or genomic data, are currently used to identify resistotypes of bacterial isolates. Metagenomic sequencing and bioinformatics analysis methods are used to identify resistome and resistotypes of microbial communities.
Conclusion. Identification of resistotypes provides additional assessment of resistome in different microbial populations. Resistotype analysis can be applied both in clinical practice, to select the most appropriate method of therapy, and in agriculture, to improve the control of antibiotic resistance of microorganisms pathogenic to animals.
Keywords
Full Text
Intensive use of antimicrobials leads to an increase in the number of resistance genes both among microorganisms associated with human body biotopes and among microorganisms in the human environment. The latter may include microorganisms inhabiting the biotopes of farm and domestic animals, as well as those in the soil or on the surfaces of objects surrounding humans. According to the World Health Organization, about 50% of antimicrobials used are not prescribed by a doctor1. However, even if a person is not using antibacterial drugs, they may still be exposed to environmental bacteria, which can lead to the spread of antimicrobial resistance (AMR) genes in their own microflora. AMR genes can be acquired not only by the commensal microflora of the human body, but also by pathogenic or opportunistic microorganisms, which can significantly complicate the treatment of infectious diseases.
The totality of all AMR genes in one biotope is called the resistome. The resistome of human biotopes is formed mainly by the commensal microflora, and resistance genes of pathogenic microorganisms constitute only a small part of it [1].
The use of full-genome metagenomic sequencing makes it possible to identify the majority of AMR genes that make up the resistome, including AMR genes in the genomes of uncultivated microorganisms. This approach is currently used to objectively characterize the resistome and subsequently identify resistotypes. The term “resistotype” can be met more frequently in scientific literature. Resistotypes are usually referred to as specific clusters within a resistome that possess a characteristic phenotypic profile or a set of antimicrobial resistance genes [2, 3].
Identification of resistotypes is not yet a common practice for microbiological studies related to resistance analysis. Upon examination of the statistics in the PubMed database, the query “resistotype*[text]” shows 43 publications in the last 10 years in which this term appears. For comparison, the query “antibiotic resistance gene*[text]” found 8848 articles (query date: 06.08.2024). At the same time, studies in which resistotypes are identified and characterized are predominantly published in highly ranked journals [4–7].
The aims and objectives of this review are to analyze and summarize current data on the methods of identification and analysis of resistotypes of individual microorganism species and bacterial communities.
Literature sources were searched using the PubMed database using the query “resistotypes[text] OR resistotype[text]” (query date 06.08.2024). Full-text articles published between 2014 and 2024 were reviewed. Relevant articles were also searched in the Google Scholar database using the query “resistotype” for the same period. Articles describing individual resistotypes of bacteria or microbial communities were included in the study. Papers for which it was not possible to obtain the full text of the article, as well as articles written in languages other than English, were excluded from the search results (no Russian-language publications on this topic were found, including in the eLIBRARY database). Forty-three articles were found in the PubMed database for the above-mentioned query. Also, 5 articles were additionally included in the review based on the search results in the Google Scholar database (Fig. 1). Two publications in Turkish and 22 articles lacking descriptions of individual resistotypes were excluded from the analysis.
Fig. 1. Scheme for selecting publications for review.
A total of 24 publications were selected based on the inclusion/exclusion criteria presented in this review. All articles analyzed described resistotypes of individual bacterial isolates or metagenomic communities from both human and farm animals. All publications used phenotypic research methods to determine resistotypes of bacterial isolates, rarely supplemented by genetic or genomic data. Metagenomic sequencing and bioinformatic analysis methods were used to determine the resistome and resistotypes of microbial communities (Table 1).
Table 1. Distribution of publications by object of study and methods used to determine resistotypes
Type of research | Object | |
human | farm animals | |
Metagenomic studies | 6 | 1 |
Phenotypic methods/isolates | 11 | 6 |
Resistotypes of selected bacterial species
Studies describing the resistotypes of individual bacterial species are currently devoted either to the analysis of clinical isolates isolated from samples of patients with various infectious diseases or to the analysis of bacterial isolates isolated from samples of diseased farm animals. Both the classical disc diffusion test for determining phenotypic sensitivity to antibiotics and methods of genetic testing and whole-genome sequencing are used to determine resistotypes in individual bacterial species. The publications considered in this section of the review can be conditionally divided into two groups:
Studies describing resistotypes of bacteria clinically relevant to humans [2–4, 7–20];
Studies describing the resistotypes of pathogens of infectious diseases of farm animals [21–27].
Resistotypes of clinically relevant bacteria
The traditional approach to the determination of resistotypes of individual microorganism species is the use of phenotypic methods of antibiotic susceptibility testing, with the vast majority of studies devoted to the study of microorganisms belonging to the ESKAPE group of pathogens. In particular, a study by B. Pérez-Viso et al. presented the results of identification of resistotypes of bacterial isolates of Escherichia coli (n = 9,514), Klebsiella pneumoniae (n = 2,137) and Enterobacter cloacae (n = 516) on the basis of disk-diffusion sensitivity testing to ampicillin, cephalothin, cefoxitin, amoxicillin and clavulanic acid [8]. The study revealed 5 resistance variants of bacterial isolates, of which 4 variants were found in all 3 bacterial species (Table 2).
Table 2. Characteristics of resistotypes from the publication B. Pérez-Viso et al. [7]
Resistotype designation in the text of the publication | Features | Antibiotic sensitivity | Antibiotic resistance | Association with carriers, if specified | Cohort size |
Wild-type | – | All tested | – | E. coli, K. pneumoniae, E. cloacae | 7101 |
BSBL resistotype | Presence of broad-spectrum beta-lactamases, absence of extended-spectrum β-lactamases | Cephalothin, cefoxitin | Ampicillin | E. coli | 3653 |
ESBL resistotype | Presence of extended-spectrum β-lactamases, absence of β-lactamase ampC and carbapenemase | Cephalosporins | E. coli, K. pneumoniae, E. cloacae | 1217 | |
AmpC resistotype | Presence of ampC β-lactamase, absence of carbapenemase and extended-spectrum β-lactamases | Cephalosporins and clavulanic acid | E. coli, K. pneumoniae | 195 |
Some of the isolates analyzed did not fall into any of the described 4 groups. The resistance spectra of these isolates were different for different bacterial species.
The relative prevalence of resistotypes within the same species of bacterial isolates was different for different species. If in E. coli isolates wild-type and BSBL-type resistotypes were most common (49 and 37%, respectively), then in K. pneumoniae and E. cloacae isolates the wild-type resistotype and a group of unclassified resistotypes prevailed. At the same time, the BSBL resistotype was identified only in E. coli isolates. Although this publication provides a detailed characterization of the resistotypes of the isolates, a significant drawback is the lack of an analysis comparing resistotypes with the metadata of the patients from whose specimens the isolates were isolated [8].
Among other publications devoted to the analysis of resistotypes of individual bacteria, there are works similarly describing the resistotypes of K. pneumoniae [12, 15–17], Pseudomonas aeruginosa [11, 18], Acinetobacter spp. [11, 13, 15, 17], as well as Enterococcus spp., Staphylococcus aureus, Enterobacter spp. [15] and Stenotrophomonas maltophilia [14]. In all the above-mentioned publications, the establishment of bacterial resistance was performed by testing their sensitivity to antibiotics using the disk diffusion method or E-tests.
Recently, a study demonstrating the variability of resistance patterns of Burkholderia cenocepacia isolates isolated from the sputum of a patient with cystic fibrosis has been published [9]. Sputum samples were collected from 1 patient for 63 months (more than 5 years), from which B. cenocepacia were isolated. The obtained isolates were tested for sensitivity to ceftazidime, meropenem, minocycline, and trimethoprim-sulfamethoxazole. A total of 11 isolates were analyzed, each with a unique resistotype (Fig. 2). In this article, the term “resistotype” was used to refer to a specific AMR profile characteristic of an isolate, and resistotypes can change over time. While an isolate isolated at the beginning of the study was sensitive to all 4 antibiotics, the resistance profile of isolates isolated in subsequent months changed, and the observed changes were not always associated with an increase in resistance. An isolate resistant to all 4 antibiotics was obtained only at the 18th month of the study and was not detected in the following months.
Fig. 2. Description of 11 resistotypes of B. cenocepacia isolates from adult cystic fibrosis patient samples over a period of 63 months.
Resistotype profiles were based on the sensitivity of the isolates to 4 antibiotics (ceftazidime, meropenem, minocycline, and trimethoprim-sulfamethoxazole). S, sensitive; I, intermediately resistant; R, resistant. CAZ, ceftazidime; COTRIM, trimethoprim-sulfamethoxazole; MERO, meropenem; MINO, minocycline. Data are based on the publication [9].
The most characteristic study describing the establishment of bacterial resistance by combining phenotypic and genetic testing is the publication [7] devoted to the study of microbiological and clinical characteristics of bacteria of the Serratia genus, which presents the results of work conducted over 16 years (2005–2020). Serratia are opportunistic pathogenic microorganisms that cause purulent-inflammatory diseases of various localizations in humans. The most studied species of this genus is S. marcescens. This pathogen is the etiologic agent of diarrheal diseases, meningitis, arthritis, sepsis and urinary tract infections. Species of the Serratia genus are considered one of the main sources of bacteremia in perinatal centers. The Serratia spp. isolates included in the cited study were isolated from clinical samples of patients at the Ramón y Cojal University Hospital (Madrid), whose ages ranged from 21 days to 97 years. A total of 107 isolates were selected for whole-genome sequencing and subsequent analysis of Serratia spp. resistome.
As a result of bioinformatic analysis, 4 resistotypes were identified among the analyzed sample of isolates (Table 3).
Table 3. Characteristics of Serratia spp. resistotypes from article [7]
Resistotype designation in the text of the publication | Major driver genes of resistotype | Antibiotic resistance | Association with carriers | Cohort size |
Resistotype 1 | aac6-Ic, blaSST-1, tet41 | Aminoglycosides, β-lactams, tetracycline | S. marcescens, S. nematodiphila, S. ureilytica | 62 |
Resistotype 2 | aac6-Ic, blaSRT-1, blaSHV-12, blaVIM-1, cmlB1 | Aminoglycosides, β-lactams, carbapenems, cephalosporins, chloramphenicol | 37 | |
Resistotype 3 | aac6-Ic, aadA1, blaSRT-1, blaVIM-1, tet41, sul1, catA1, catB2, dfrB1, mphE, msrE | Aminoglycosides, β-lactams, carbapenems, tetracyclines, macrolides | 6 | |
Resistotype 4 | aac6-Ic, blaSRT-1, blaSHV-12, blaLAP-2, qnr-S1 | Aminoglycosides, β-lactams, carbapenems, fluoroquinolones | 2 |
Resistotype I (aac6-Ic, blaSST-1 and tet41 genes) and resistotype II (aac6-Ic and blaSRT-1) were represented most frequently (58 and 34.6%, respectively). Resistotype III was detected in 6 isolates producing carbapenemases (aac6-Ic, aadA1, blaSRT-1, blaVIM-1, tet41, sul1, catA1, catB2, dfrB1, mphE, msrE), and resistotype IV was represented by 2 strains producing blaSHV-12 BLRS (aac6-Ic, blaSRT-1, blaSHV-12, blaLAP-2, and qnr-S1).
The bla SST-1 and bla SRT-1 genes encoding inducible β-lactamase ampC genomes of S. marcescens were detected in all isolates. Their phenotypic expression was observed in 89.7% of the tested isolates susceptible to cefotaxime (87 out of 97 isolates). The cited study provides a detailed description of the identified Serratia spp. resistotypes, but does not compare the resistotypes with the metadata of the patients from whose specimens the analyzed isolates were isolated [7].
Resistotypes of infectious agents of farm animals
Resistome of pathogens of infectious diseases of animals is a subject of active study by researchers, including due to close human contact with farm animals. For example, S. Chhabra et al. described resistotypes of Rhodococcus equi isolates from samples of foals with respiratory diseases [21]. This pathogen is present in soil as well as in the intestinal tract of cattle, horses, sheep, pigs and several other animals. It can cause respiratory disease in foals aged 1–4 months. R. equi is considered a serious threat to the horse industry due to the high morbidity and mortality in infected young horses. In the study mentioned above, 28 clinical isolates of R. equi obtained from samples of foals with respiratory diseases from different parts of the states of Haryana and Rajasthan in India were investigated. The collected isolates were screened for resistance to 33 antibiotics by disk diffusion method. All isolates showed similar resistance results to 29 antibiotics. Differences in resistance profiles were observed only in relation to 4 antimicrobials: amoxicillin, gentamicin, colistin and streptomycin. Based on these differences, 10 resistotypes (R1–R10) were identified. Resistotypes R1 (resistance to amoxicillin, gentamicin, and streptomycin; sensitivity to colistin) and R4 (resistance to gentamicin and streptomycin; sensitivity to amoxicillin and colistin) were the most frequent. The article emphasizes that the study of resistotypes can help in determining the source of infection and spread of disease, and can be used to select effective therapy and control disease in a particular geographic area or on a particular farm.
Similar work was carried out with Enterococcus spp. isolates obtained from bird samples from 40 poultry farms in Serbia [22]. The sensitivity of the isolates to antibiotics was also determined by disk diffusion method. The study identified different species of the genus Enterococcus with different resistance profiles, including E. faecalis, E. faecium, E. hirae, E. durans and E. thialandicus. Meanwhile, E. faecalis and E. faecium were most associated with multidrug resistance (MDR) to antibiotics. In this study, 18 resistotypes associated with resistance to different classes of antibiotics were identified. For example, resistotypes characterized by resistance to tetracycline and doxycycline alone were identified, while other resistotypes were characterized by MDR to several classes of antibiotics. The identified MDR strains may pose a risk to both avian and human health as they may contribute to the spread of antibiotic resistance through the food chain.
A large study by J. Alvarez et al. analyzed the resistotypes of 3047 Salmonella isolates from the feces of farm birds (laying hens, broiler chickens and turkeys) in Spain [25]. Sensitivity to 9 antibiotics (ampicillin, ciprofloxacin, nalidixic acid, chloramphenicol, gentamicin, sulfamethoxazole, tetracycline, trimethoprim and colistin) was determined using serial dilution method. The study identified 94 resistotypes, of which 33 were found in all bird species, 19 were characteristic only for broiler chickens, 22 for turkeys, and 3 for laying hens.
Similar studies describing the resistotypes of bacterial pathogens of farm animals were also conducted on poultry farms in Zimbabwe (for Salmonella enteritidis isolates [23]) and Italy (for Campylobacter jejuni isolates [24]). Also, a study by D. Cid et al. described the resistotypes of Pasteurella multocida isolates from pig and sheep samples [26].
Thus, in the reviewed publications, the term “resistotype” is understood as a certain phenotype of resistance of microorganisms to a set of antibiotics, detected by disk diffusion method, serial dilution method or E-test strip method.
Resistotypes of commensal microbial communities
The microbiome associated with the host organism (human or animal) is a complex and dynamic community of multiple species of microorganisms: bacteria, archaea, viruses and fungi inhabiting different biotopes, mainly skin, mucous membranes and gastrointestinal tract. The microbiome plays an important role in maintaining the health of the host organism and may be associated with a number of diseases [28]. Drugs affecting the host also have an effect on the microbiome, with the most pronounced effect observed for antimicrobials [29, 30].
Even a healthy person who does not take antibiotics is constantly in contact with antibiotic-resistant microorganisms, which can be carried by other people or pets. This leads to the possibility that AMR genes may enter his or her own microbiota, becoming part of the individual resistome. Such events are most likely to occur in populations with a high frequency of antibiotic consumption [31].
In determining resistomes and resistotypes of microbial communities, genetic testing methods are used exclusively, in contrast to works on the study of resistotypes of individual bacterial species. Of the genetic methods used for resistome analysis, metagenomic sequencing followed by bioinformatic analysis is the most informative.
The first mention of the term “resistotype” in relation to microbial communities associated with the human body dates back to a 2013 study [3]. In this study, gut microbiota resistotypes were analyzed based on metagenomic data from 275 healthy volunteers from America, Denmark, Spain, France, Italy, Japan, China and India. The analysis revealed the presence of potential resistance genes to 53 different antibiotics. The sample included adults without serious pathologies (267 samples), children (4 samples) and infants (4 samples), allowing comparative analysis of resistotypes of different age groups.
AMR gene prevalence data were obtained for each metagenome. Genes occurring in less than 10% of metagenomes were excluded from the analysis. Between Class Analysis (a modification of Principal component analysis, which is more robust to outliers and data noise) was performed for the obtained representation profiles. The study identified 4 resistotypes by cluster analysis (Table 4).
Table 4. Characteristics of human bacterial community resistotypes
Object of study | Resistotype designation in the text of the publication | Resistotypes (main drivers, if specified) | Antibiotic resistance | Associations with bacteria | Cohort size | Resistotype properties | Reference |
Nasopharyngeal microbiome of preterm infants | R1 | patA, patB, rlmA(II) | Fluoroquinolones | Streptococcus mitis/oralis, Gemella haemolysans/sanguinis | 66 | One of the most represented resistotypes in premature infants | [18] |
R2 | SST-1, AAC(6')-Ic, tet(41), mexl | β-Lactams, aminoglycosides, tetracycline, multidrug resistance (MDR) | S. marcescens/ nematodiphila | One of the most represented resistotypes in premature infants. It persists for a long time | |||
R3 | blaZ | β-Lactams | S. aureus, S. epidermidis | ||||
The airway microbiome in bronchiectasis | RT1 | hmnM, PatA, tetB(46), ErmX | Fluoroquinolones, tetracyclines, MDR | Haemophilus influenzae, Rothia mucilaginosa, Streptococcus spp. | 280 | RT1 is a more clinically favorable resistome profile showing less resistance gene diversity | [2] |
RT2 | mexM, basA, PA_catB7, bcr_1, APH(3)''Ib | Aminoglycoside, phenicol, bicyclomycin, MDR | P. aeruginosa, K. pneumoniae | RT2 is associated with more exacerbations, worse lung function, and greater disease severity | |||
The gut microbiome of healthy individuals | Resistotype 1A | Bacitracin, vancomycin, tetracycline, F3H8F5 (lincosamine, streptogramin B and macrolides) | 267 | Predominantly sampled from Europe and Japan | [3] | ||
Resistotype 1B | Vancomycin, bacitracin, tetracyclines, cephalosporins | Predominantly sampled from Europe and the USA | |||||
Resistotype 1C | Fosmidomycin, cephalosporin | Predominantly sampled from the US | |||||
Resistotype 2 | Tetracycline, F3H8F5 (lincosamine, streptogramin B and macrolides), cephalosporin, lincomycin, macrolide, J3I4 and trimethoprim | Predominantly Chinese sample | |||||
The gut microbiome of healthy individuals | Background | Glycopeptides, tetracyclines | Coprococcus eutactus, Eubacterium siraeum | 3034 | Associated with non-pathogenic microorganisms | [4] | |
FAMP | Fluoroquinolones, MDR, polypeptides, fluoroquinolones, fosfomycin, aminoglycosides, sulfonamide | E. coli, Proteus mirabilis | 2338 | Associated with pathogenic microorganisms. Occurs in metagenomes of patients with colorectal cancer (51.9%), metabolic diseases (58.2%), intestinal infections associated with Shiga toxin-producing E. coli (79.4%), and cholera | |||
Microbiota of the oral cavity biofilm | Resistotype 1 | High representation of mefA, msrD, ermB, blaCSP(1). Low representation of tet32 and tetQ | 179 | Prevalent in all groups of people (predominantly in healthy individuals and those with caries) | [19] | ||
Resistotype 2 | High representation of ermF and tet32, tetQ. Low representation of ermB and blaCSP(1) | Present in all groups of people (predominantly in healthy individuals and those with caries) | |||||
Resistotype 3 | High representation of pgpB. Low representation of mefA, msrD, ermF, ermB | Present only in samples of people with periodontitis | |||||
Intestinal microbiota of healthy individuals | 6 resistotypes were identified | Resistotype 1 is ANT, Resistotype 3 Tet(M) and class C β-lactamases. Resistotype 4 by Tet(X) and class A β-lactamases. Resistotype 6 by β-lactamases of class B1, Sul | Resistotypes 1 and 3 — bacterial order Clostridiales. Resistotype 4 — bacterial genus Bacteroides. Resistotype 6 — genus Prevotella | 663 | The first 4 resistotypes were the most represented, each comprising about 20% of the samples studied. The fifth and sixth resistotypes represented 8.7 and 7.5%, respectively | [20] |
Resistotype 1 included most of the gut microbiota samples studied. The authors categorized this resistotype into 3 subtypes: 1A, 1B and 1C. Resistotype 1A was the most common among European and Japanese study participants. Resistotype 1B included samples from different regions; no pronounced geographic specificity was observed for this subtype. Resistotype 1C included a significant number of samples from the USA (about 60% of American microbiomes). This resistotype was characterized by higher resistance to fosmidomycin and cephalosporins, which may be associated with the specifics of the use of these antibiotics in the USA.
Resistotype 2 was specific to the Chinese population (24 out of 30 Chinese samples) and was also characterized by a high content of tetracycline, lincomycin and macrolide resistance genes. This reflects the peculiarities of antibiotic use in China, where there is a higher use of antibiotics compared to other countries [32].
The MetaHIT study found significant geographical differences in the number of identifiable AMR genes [33]. In particular, a higher number of AMR genes were found in the intestines of people from Southern Europe compared to people from Northern Europe and the USA. A higher number of AMR genes was detected in children from Japan compared to children from India, but a correction should be made for the small sample size: there were only 2 pediatric samples from India and Japan each. It is worth noting that infants from Japan (4 samples) had a high number of AMR genes detected.
A study by E. Ruppé et al. was performed using metagenomic data obtained within the MetaHIT consortium [33], the DMM (Dirichlet-Multinomial Mixture Models) method was used to identify resistotypes using the Laplace criterion to determine the optimal number of clusters [20]. We analyzed 663 metagenomic samples, which we were able to cluster into 6 resistotypes. Among them, the first 4 resistotypes were the most common, each comprising about 20% of the samples analyzed. The fifth and sixth resistotypes were present in 8.7 and 7.5% of the samples, respectively. Resistotype 1 was enriched in ANT aminoglycoside resistance genes, while resistotype 3 was dominated by tet(M) tetracycline resistance genes and class C β-lactamase resistance genes. Resistotype 4 was enriched with tet(X) tetracycline resistance genes and class A β-lactamases, while resistotype 6 was enriched with class B1 β-lactamases and sul sulfonamide resistance genes. Resistotypes 1 and 3 had a higher diversity of AMR genes and were associated with the Clostridiales bacterial order. Resistotype 4 was associated with bacteria of the Bacteroides genus (they contain tet(X) and β-lactamase class A genes), and resistotype 6 — with bacteria of the Prevotella genus.
Over the past 2 years, several papers have appeared in highly ranked journals developing the topic of characterization of resistotypes of microbial communities inhabiting natural human biotopes. Perhaps such interest in the patterns of AMR gene distribution is due to public concern about the consequences of the COVID-19 pandemic associated with excessive consumption of antibiotics2.
In particular, K. Lee et al. attempted to assess the impact of antibiotics use on the human microbiome at the population level [4]. The study included metagenomic data obtained for the microbiota of different biotopes of the human body: oral cavity, skin, upper respiratory tract, vagina, but the main array (n = 5372) of samples consisted of metagenomes of the intestinal microbiota. Metagenomes from different countries were included in the study: Austria, Canada, China, Denmark, Denmark, Spain, France, Germany, Israel, Italy, Kazakhstan, Madagascar, Netherlands, Spain, Sweden and USA. Metagenomes from healthy volunteers accounted for 3,565 of the 8,972 samples analyzed. Using this sample, the authors showed that the representation of AMR genes correlates with the level of antibiotic consumption per capita in the analyzed country, and this correlation is primarily observed for AMR genes adjacent to mobile genetic elements.
In analyzing the above-mentioned samples, 422 AMR gene families were identified using the CARD (Comprehensive Antibiotic Resistance Database). The resistome profiles obtained for the metagenomic samples were clustered by NMDS (non-metric multidimensional scaling) using Bray-Curtis distances, resulting in the identification of 2 resistome clusters. The separation into 2 resistotypes was confirmed using PAM (partitioning around medoids), UMAP (Uniform Manifold Approximation and Projection) and k-means methods.
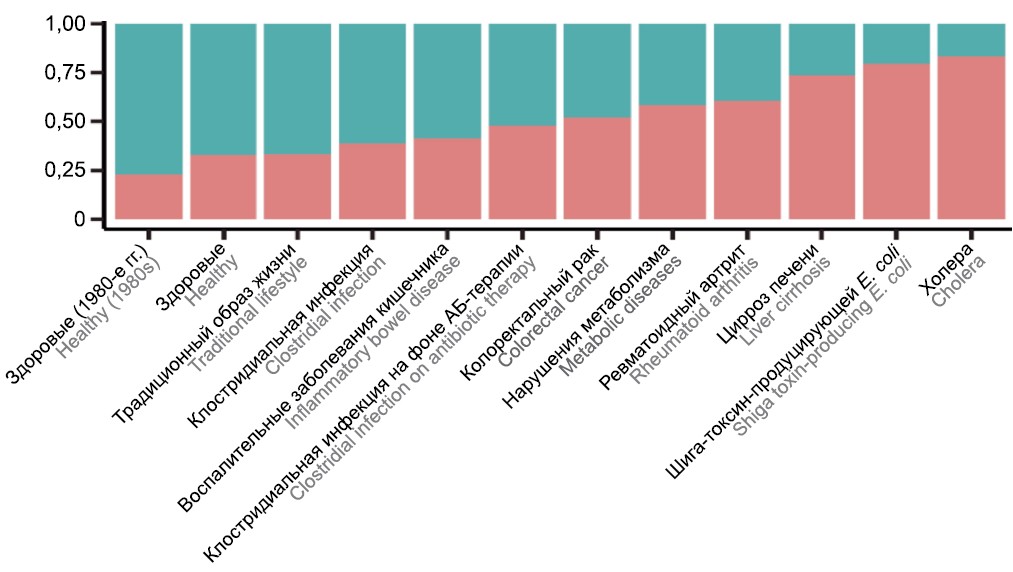
When analyzing the frequencies of occurrence of resistotypes and the representation of AMR genes in them, it was found that the resistotype with a lower frequency of occurrence in the population is more than 10 times enriched with resistance genes to fluoroquinolones, fosfomycins, aminoglycosides and peptide antibiotics, as well as genes determining MDR, compared to the other resistotype, which is found in a larger number of samples. The first, rarer resistotype was designated by the abbreviation FAMP, based on the first letters of the names of the antibiotic groups whose resistance genes are characteristic of this resistotype (Fluoroquinolones, Fosfomycins, Aminoglycosides, Multi-drug resistance, Peptide antibiotics). The other resistotype was labeled as background. Importantly, no association with enterotypes was found for the identified resistotypes, but associations with specific bacterial taxa were established. Thus, the species associated with the FAMP resistotype were predominantly from the Proteobacteria phylum, in particular, from the Enterobacteriaceae family. At the same time, the frequency of FAMP resistotype occurrence was correlated with patient groups by health status: in healthy patients, the proportion of FAMP samples was minimal, while FAMP resistotype was most frequently found in groups of patients with intestinal infections such as cholera (83.3%) and Shiga toxin-producing Escherichia coli (79.4%) (Fig. 3).
Fig. 3. Representation of FAMP (bottom of the columns) and background (top of the columns) resistotypes in the gut microbiota of different categories of people [4].
Also, this study used longitudinal data from 12 healthy volunteers receiving a 4-day course of a mixture of 3 broad-spectrum antibiotics: meropenem, gentamicin and vancomycin [34]. By analyzing these data, it was shown that antibiotic use leads to a transition to FAMP-resistance for the majority of patients within 8 days after the end of the antibiotics course and is accompanied by an increase in the total number of AMR genes. This effect partially persists 42 days after the end of the course, but by day 180 after the end of the course of antibiotic use, patients return to the original resistotype (background). The authors attribute the observed effect to AMR genes associated with single Proteobacteria species.
The authors suggest that the use of antibiotics may lead to the emergence of resistance primarily in pathogenic and opportunistic gram-negative microorganisms that may enter the human gut during intestinal infections, subsequently transferring resistance genes to commensal bacteria. The authors also suggest that this process of sharing antibiotic resistance genes within the gut microbial community may take a long time, and the speed of this process may depend on the overall intensity of antibiotic use in the population. These hypotheses help to explain the association of the FAMP resistotype with intestinal infections and Proteobacteria as drivers of this resistotype, as well as with the overall level of antibiotic consumption in the population.
The topic of resistotype variability is raised by A. Dhariwal et al., who studied the formation of the microbiome and nasopharyngeal resistome in premature infants with the assessment of the influence of early antibiotic use on this process [18]. The study included 66 premature infants whose nasopharyngeal aspirate samples were collected during the first 6 months of life. The total sample volume amounted to 181 samples. According to the results of metagenomic analysis with subsequent data processing, 3 main resistotypes were identified in this sample, the characteristics of which are shown in Table 4.
The authors showed that early antibiotic administration had a transient effect on resistome and distribution of resistotypes [18]. Exposure to antibiotics led to an increase in the diversity and number of antibiotic-resistance genes. However, this effect was short-lived, and differences in resistotypes between groups with and without antibiotic administration smoothed out by 6 months of corrected age. The R2 resistotype associated with Serratia bacteria persisted longest after antibiotic administration.
B. Pérez-Viso et al. found an association between the airway resistotype of patients with bronchiectasis and their clinical outcomes [7]. Metagenomic sequencing data of sputum samples were used to analyze the resistome and identify resistotypes. A total of 280 individuals aged 59–73 years from the UK (Scotland), Greece, Singapore, Malaysia and Italy were included in the study. Resistotypes were determined using bioinformatic data analysis, which included spectral clustering based on Bray-Curtis distances and assessment of cluster stability using maximization of the mean silhouette coefficient. This method measures differences in resistome gene profiles between different samples and groups them into clusters based on similarity. Analysis of the resistome profiles allowed us to identify 2 resistotypes (RT1 and RT2) and to perform an association of the observed resistotypes with disease outcome. RT1 was associated with more favorable clinical outcomes and contained an increased number of resistance genes to tetracyclines, macrolides, and beta-lactam antibiotics. RT2 was associated with unfavorable clinical outcomes and was characterized by an increased number of resistance genes to aminoglycosides, chloramphenicol, bicyclomycin, peptide antibiotics, as well as an increased number of MDR determinant genes (Table 4).
In samples belonging to RT1, there was an increased relative representation of bacteria of H. influenzae and R. mucilaginosa species, as well as several species of Streptococcus genus. In samples belonging to the RT2 resistotype, an increased representation of P. aeruginosa and K. pneumoniae pathogens was observed. Successful eradication of P. aeruginosa in patients led to a switch from RT2 to RT1, accompanied by a decrease in resistance gene diversity and improved clinical outcomes. The study highlights the importance of resistome analysis in predicting clinical outcomes in patients with bronchiectasis. The authors of the paper believe that the identification of RT1 and RT2 opens new opportunities for targeting therapy and improving the clinical outcome of the disease.
A.C. Anderson et al. described the features of oral biofilm resistome in healthy patients, as well as patients with caries and periodontitis [19]. This study utilized metagenomic data obtained from samples of 179 individuals. Using hierarchical clustering on the Jaccard difference matrix, 3 resistotypes were identified. Resistotype 1 was characterized by high representation of mefA, msrD, ermB, and blaCSP(1) genes and low representation of pgpB, tet32, and tetQ genes. Resistotype 2 had high representation of ermF, tet32 and tetQ genes and low representation of ermB and blaCSP(1) genes. Resistotype 3 was characterized by high representation of pgpB genes and low representation of mefA, msrD, ermF and ermB genes. Resistotype 3 was present only in the samples of patients with periodontitis, while resistotypes 1 and 2 were present in all groups, but to the greatest extent in the groups of healthy and caries patients (Table 4).
The only study to date describing the resistotypes of farm animal microbiota has analyzed the rumen microbiota resistome of cows and evaluated the relationship between resistotypes and the nutritional value of milk [27]. In this study, 49 samples of rumen microbiota (the initial stomach compartment of ruminants) collected from cows from two different farms in China were analyzed. The study was divided into two parts. The first part, which included samples from 33 cows, was to evaluate the effect of feed intake on animal resistome and resistotypes. In the second part of the study, the protein content in milk of cows was determined depending on the rumen microbiota resistome. Sixteen animals with high and low protein content in milk were included. AMR genes were detected using full genome metagenomic sequencing, bioinformatic data processing was performed using the CARD database. Resistotypes were identified using clustering with a partitioning algorithm around the medoid. The study identified 4 resistotypes associated with milk protein production. The rumen microbiota of cows with low milk protein levels was assigned to one resistotype and characterized by a high content of AMR genes. The most represented in this group were the mfd gene, which determines resistance to fluoroquinolones, and the sav1866 gene associated with MDR. No relationship was found between resistome and consumption of different amounts of feed in the study. The authors conclude that the cow rumen microbiome and associated antibiotic resistance profiles may influence the quality of dairy products.
Discussion
The concept of resistotype represents a new characterization of bacterial communities, distinct from their taxonomic composition. In contrast to traditional approaches that focus on individual resistance genes or phenotypic manifestations of resistance, the analysis of resistotypes allows us to consider resistance at a more comprehensive level, integrating different genes and their interactions within microbial populations or entire microbial communities.
On our part, this is not the first reference to the topic of defining the role of resistome in microbial communities. Being the first scientific group in Russia to describe the features of the gut microbiota of our country’s inhabitants (healthy volunteers) [35], we also proposed a tool for resistome analysis [36] and correlated the resistome data of the gut microbiota of Russians with the then available ideas about the phenomenon of resistome of microbial communities [37].
Today we address the topic of resistotypes as a taxonomy-independent characteristic of the microbiome based on our own experience of detecting two resistotypes of the gut microbiota in patients hospitalized with COVID-19. The two resistotypes we detected differed in the number of AMR genes, with half of the patients experiencing a change in resistotype during therapy. It is worth noting that for the microbiota of the oropharyngeal tract of the same patients, we observed a single resistotype characterized by resistance genes to macrolides, fluoroquinolones, and lincosamide [38].
Studies on resistotypes use different methodological basis for their identification and characterization. Studies devoted to the determination of resistotypes of individual bacterial species are usually based on phenotypic characterization of the sensitivity of these microorganisms to different antibiotics and only rarely use genetic testing. Resistotype analysis of microbial communities, such as the human gut microbiome, is realized through whole-genome metagenomic sequencing, which allows the detection of most of the AMR genes in the resistome, including those in uncultured organisms. This method provides detailed information on the genetic composition of microbial communities and identifies specific clusters of AMR genes that form resistotypes. Thus, different interpretations of the concept of “resistotype” depending on the object of study and the way of its characterization can be found in the scientific press today. In the case of the study of individual microorganism species, a resistotype refers to a unique phenotypic susceptibility profile, whereas in the case of metagenomic analysis of microbial communities, a resistotype is the result of clustering the identified AMR genes and selecting the optimal number of clusters.
The importance of the resistotype as an independent characteristic of the microbial community is yet to be realized. In clinical practice, the identification of resistotypes in the future may play a role in the selection of rational antimicrobial therapy. Knowledge of the presence of a specific resistotype in the patient’s microbiota will allow physicians to choose the most effective combinations of antibiotic drugs, minimizing the risk of developing AMR. Thus, resistotyping can significantly accelerate and improve the decision-making process in the treatment of infectious diseases.
In the agricultural sector, the main objective of the study of resistotypes is to identify the resistance spectrum of microorganisms pathogenic to animals. This is particularly relevant for determining the risks of transferring resistance genes from animals to humans through the food chain. For example, identifying resistotypes in pathogens such as R. equi in horses or Enterococcus spp. in birds helps to assess the potential health consequences for both the animals themselves and the humans who come into contact with these animals, as well as to develop effective methods for controlling the spread of these pathogens.
Understanding the characteristics of animal microbiota resistotypes can help to improve agricultural production performance. For example, in a study of the relationship between rumen microbiota resistotypes and the quality of dairy products, it was shown that certain resistotypes can be associated with the protein content of milk. Thus, monitoring and control of resistotypes can be used to improve the efficiency of production and ensure the safety of agricultural products.
Conclusion
Resistotypes represent a new characteristic of bacterial communities, considered separately from taxonomic composition. Identification of resistotypes allows an additional assessment of resistomes in different microbial populations. Various factors may play a role in the formation of resistotypes of the microbiota of individuals: clinical, cultural, geographical, etc., which emphasizes the necessity to take into account regional characteristics when developing strategies to combat AMR. The identification of resistotypes holds promise both in clinical practice, where it may facilitate the selection of the most appropriate therapy, and in agriculture, where this approach can be used to improve the control of AMR of microorganisms pathogenic to animals.
1 The abuse of antibiotics leads to an increase in mortality from infections. URL: https://news.un.org/ru/story/2019/11/1367331 (In Russ.)
2 WHO reports widespread overuse of antibiotics in patients hospitalized with COVID-19. URL: https://www.who.int/news/item/26-04-2024-who-reports-widespread-overuse-of-antibiotics-in-patients--hospitalized-with-covid-19
About the authors
Alexander V. Pavlenko
Research Institute for Systems Biology and Medicine
Author for correspondence.
Email: pavav@mail.ru
ORCID iD: 0000-0002-9549-0289
researcher, Laboratory of mathematical biology and bioinformatics
Russian Federation, MoscowIgnat V. Sonets
Research Institute for Systems Biology and Medicine
Email: ignatsonets@gmail.com
ORCID iD: 0009-0009-7266-6657
junior researcher, Laboratory of mathematical biology and bioinformatics
Russian Federation, MoscowAleksandr I. Manolov
Research Institute for Systems Biology and Medicine
Email: paraslonic@gmail.com
ORCID iD: 0000-0003-3912-429X
Cand. Sci. (Biol.), leading researcher, Head, Laboratory of digital monitoring of human health
Russian Federation, MoscowElizaveta V. Starikova
Research Institute for Systems Biology and Medicine
Email: e.starikova@sysbiomed.ru
ORCID iD: 0000-0001-6582-210X
researcher, Laboratory of mathematical biology and bioinformatics
Russian Federation, MoscowElena N. Ilina
Research Institute for Systems Biology and Medicine
Email: ilinaen@sysbiomed.ru
ORCID iD: 0000-0003-0130-5079
D. Sci. (Biol.), Сorresponding member of RAS, chief scientist, Head, Laboratory for mathematical biology and bioinformatics
Russian Federation, MoscowReferences
- O'Connor L., Heyderman R. The challenges of defining the human nasopharyngeal resistome. Trends Microbiol. 2023;31(8):816–31. DOI: https://doi.org/10.1016/j.tim.2023.02.008
- Mac Aogáin M., Ivan F.X., Jaggi T.K., et al. Airway "resistotypes" and clinical outcomes in bronchiectasis. Am. J. Respir. Crit. Care Med. 2024;210(1):47–62. DOI: https://doi.org/10.1164/rccm.202306-1059oc
- Ghosh T.S., Gupta S.S., Nair G.B., Mande S.S. In silico analysis of antibiotic resistance genes in the gut microflora of individuals from diverse geographies and age-groups. PLoS One. 2013; 8(12):e83823. DOI: https://doi.org/10.1371/journal.pone.0083823
- Lee K., Raguideau S., Sirén K., et al. Population-level impacts of antibiotic usage on the human gut microbiome. Nat. Commun. 2023;14(1):1191. DOI: https://doi.org/10.1038/s41467-023-36633-7
- Johannessen H., Anthonisen I.L., Zecic N., et al. Characterization and fitness cost of Tn7100, a novel integrative and conjugative element conferring multidrug resistance in haemophilus influenzae. Front. Microbiol. 2022;13:945411. DOI: https://doi.org/10.3389/fmicb.2022.945411
- Cawez F., Mercuri P.S., Morales-Yãnez F.J., et al. Development of nanobodies as theranostic agents against CMY-2-like class C β-lactamases. Antimicrob. Agents Chemother. 2023;67(4): e0149922. DOI: https://doi.org/10.1128/aac.01499-22
- Pérez-Viso B., Hernández-García M., Rodríguez C.M., et al. A long-term survey of Serratia spp. bloodstream infections revealed an increase of antimicrobial resistance involving adult population. Microbiol. Spectr. 2024;12(2):e0276223. DOI: https://doi.org/10.1128/spectrum.02762-23
- Moore J.E., McCaughan J., Rendall J.C., Millar B.C. Case report: the conundrum of what to pick? Antibiotic susceptibility variability in Burkholderia cenocepacia in cystic fibrosis: implications for antibiotic susceptibility testing and treatment. Br. J. Biomed. Sci. 2024;81:12749. DOI: https://doi.org/10.3389/bjbs.2024.12749
- Gajdács M., Baráth Z., Kárpáti K., et al. no correlation between biofilm formation, virulence factors, and antibiotic resistance in pseudomonas aeruginosa: results from a laboratory-based in vitro study. Antibiotics (Basel). 2021;10(9):1134. DOI: https://doi.org/10.3390/antibiotics10091134
- Çöl A., Dedeić-Ljubović A., Salimović-Bešić I., Hukic M. Antibiotic resistance profiles and genetic similarities within a new generation of carbapenem-resistant Аcinetobacter calcoaceticus–A. baumannii complex resistotypes in Bosnia and Herzegovina. Microb. Drug Resist. 2016;22(8):655–61. DOI: https://doi.org/10.1089/mdr.2015.0257
- Granov D., Dedeić-Ljubović A., Salimović-Bešić I. Characterization of carbapenemase-producing Klebsiella pneumoniae in Clinical Center University of Sarajevo, Bosnia and Herzegovina. Microb. Drug Resist. 2020;26(9):1038–45. DOI: https://doi.org/10.1089/mdr.2019.0188
- Jakovac S., Goić-Barišić I., Pirija M., et al. Molecular characterization and survival of carbapenem-resistant Аcinetobacter baumannii isolated from hospitalized patients in Mostar, Bosnia and Herzegovina. Microb. Drug Resist. 2021;27(3):383–90. DOI: https://doi.org/10.1089/mdr.2020.0163
- Gajdács M., Urbán E. A 10-year single-center experience on Stenotrophomonas maltophilia resistotyping in Szeged, Hungary. Eur. J. Microbiol. Immunol. (Bp). 2020;10(2):91–7. DOI: https://doi.org/10.1556/1886.2020.00006
- Phoon H.Y.P., Hussin H., Hussain B.M., et al. Distribution, genetic diversity and antimicrobial resistance of clinically important bacteria from the environment of a tertiary hospital in Malaysia. J. Glob. Antimicrob Resist. 2018;14:132–40. DOI: https://doi.org/10.1016/j.jgar.2018.02.022
- Mohamed E.R., Aly S.A., Halby H.M., et al. Epidemiological typing of multidrug-resistant Klebsiella pneumoniae, which causes paediatric ventilator-associated pneumonia in Egypt. J. Med. Microbiol. 2017;66(5):628–34. DOI: https://doi.org/10.1099/jmm.0.000473
- Shokri D., Rabbani Khorasgani M., Fatemi S.M., Soleimani-Delfan A. Resistotyping, phenotyping and genotyping of New Delhi metallo-β-lactamase (NDM) among Gram-negative bacilli from Iranian patients. J. Med. Microbiol. 2017;66(4):402–11. DOI: https://doi.org/10.1099/jmm.0.000444
- Dhariwal A., Rajar P., Salvadori G., et al. Prolonged hospitalization signature and early antibiotic effects on the nasopharyngeal resistome in preterm infants. Nat. Commun. 2024;15(1): 6024. DOI: https://doi.org/10.1038/s41467-024-50433-7
- Anderson A.C., von Ohle C., Frese C., et al. The oral microbiota is a reservoir for antimicrobial resistance: resistome and phenotypic resistance characteristics of oral biofilm in health, caries, and periodontitis. Ann. Clin. Microbiol. Antimicrob. 2023;22(1):37. DOI: https://doi.org/10.1186/s12941-023-00585-z
- Ruppé E., Ghozlane A., Tap J., et al. Prediction of the intestinal resistome by a three-dimensional structure-based method. Nat. Microbiol. 2019;4(1):112–23. DOI: https://doi.org/10.1038/s41564-018-0292-6
- Chhabra S., Khurana S., Kapoor P., et al. Resistotypes of Rhodococcus equi isolated from foals with respiratory problems. J. Exp. Biol. 2016;4(3S):242–8. DOI: https://doi.org/10.18006/2016.4(3S).242.248
- Velhner M., Prunić B., Aleksić N., et al. antimicrobial resistance of enterococcus isolates from poultry farms in the Republic of Serbia (Autonomous province of Vojvodina). Microorganisms. 2024;12(7):1483. DOI: https://doi.org/10.3390/microorganisms12071483
- Khumalo J., Saidi B., Mbanga J. Evolution of antimicrobial resistance of Salmonella enteritidis (1972–2005). Onderstepoort J. Vet. Res. 2014;81(1):e1–6. DOI: https://doi.org/10.4102/ojvr.v81i1.807
- Manfreda G., Parisi A., De Cesare A., et al. Typing of Campylobacter jejuni isolated from Turkey by genotypic methods, antimicrobial susceptibility, and virulence gene patterns: a retrospective study. Foodborne Pathog. Dis. 2016;13(2):93–100. DOI: https://doi.org/10.1089/fpd.2015.2048
- Alvarez J., Lopez G., Muellner P., et al. Identifying emerging trends in antimicrobial resistance using Salmonella surveillance data in poultry in Spain. Transbound. Emerg. Dis. 2020;67(1):250–62. DOI: https://doi.org/10.1111/tbed.13346
- Cid D., Fernández-Garayzábal J.F., Pinto C., et al. Antimicrobial susceptibility of Pasteurella multocida isolated from sheep and pigs in Spain – short communication. Acta Vet. Hung. 2019;67(4):489–98. DOI: https://doi.org/10.1556/004.2019.048
- Xue M.Y., Xie Y.Y., Zhong Y.F., et al. Ruminal resistome of dairy cattle is individualized and the resistotypes are associated with milking traits. Anim. Microbiome. 2021;3(1):18. DOI: https://doi.org/10.1186/s42523-021-00081-9
- Chen Y., Zhou J., Wang L. Role and mechanism of gut microbiota in human disease. Front. Cell Infect. Microbiol. 2021;11:625913. DOI: https://doi.org/10.3389/fcimb.2021.625913
- Vich Vila A., Collij V., Sanna S., et al. Impact of commonly used drugs on the composition and metabolic function of the gut microbiota. Nat. Commun. 2020;11(1):362. DOI: https://doi.org/10.1038/s41467-019-14177-z
- Panda S., El khader I., Casellas F., et al. Short-term effect of antibiotics on human gut microbiota. PLoS One. 2014;9(4):e95476. DOI: https://doi.org/10.1371/journal.pone.0095476
- Konstantinidis T., Tsigalou C., Karvelas A., et al. Effects of antibiotics upon the gut microbiome: a review of the literature. Biomedicines. 2020;8(11):502. DOI: https://doi.org/10.3390/biomedicines8110502
- Hu Y.J., Cowling B.J. Reducing antibiotic use in livestock, China. Bull. World Health Organ. 2020;98(5):360–1. DOI: https://doi.org/10.2471/blt.19.243501
- Ehrlich S.D. MetaHIT: The European Union Project on Metagenomics of the Human Intestinal Tract. In: Nelson K.E., ed. Metagenomics of the Human Body. New York;2011:307–16. DOI: https://doi.org/10.1007/978-1-4419-7089-3_15
- Palleja A., Mikkelsen K.H., Forslund S.K., et al. Recovery of gut microbiota of healthy adults following antibiotic exposure. Nat. Microbiol. 2018;3(11):1255–65. DOI: https://doi.org/10.1038/s41564-018-0257-9
- Tyakht A.V., Kostryukova E.S., Popenko A.S., et al. Human gut microbiota community structures in urban and rural populations in Russia. Nat. Commun. 2013;4:2469. DOI: https://doi.org/10.1038/ncomms3469
- Yarygin K.S., Kovarsky B.A., Bibikova T.S., et al. ResistoMap-online visualization of human gut microbiota antibiotic resistome. Bioinformatics. 2017;33(14):2205–6. DOI: https://doi.org/10.1093/bioinformatics/btx134
- Ильина Е.Н., Олехнович Е.И., Павленко А.В. Резистом микробиоты кишечника как источник формирования лекарственной устойчивости возбудителей инфекционных заболеваний человека. Патогенез. 2017;15(3):20–32. Ilyina E.N., Olekhnovich E.I., Pavlenko A.V. The gut microbiota resistome provides development of drug resistance in causative agents of human infectious diseases. Pathogenesis. 2017;15(3):20–32. DOI: https://doi.org/10.25557/GM.2017.3.8494 EDN: https://elibrary.ru/zwohrj (in Russian)
- Starikova E.V., Galeeva Y.S., Fedorov D.E., et al. Oropharyngeal resistome remains stable during COVID-19 therapy, while fecal resistome shifts toward a less diverse resistotype. iScience. 2024; 27(12):111319. DOI: https://doi.org/10.1016/j.isci.2024.111319
Supplementary files